对组装结果中GAP左右批量设计引物
对组装结果中GAP左右批量设计引物
对于组装结果,常常会有一些GAP,我们需要用一代测序去填洞。这样就需要我们提取GAP左右的序列设计引物,这样可以用PCR把GAP扩增出来然后进行一代测序。往往GAP很多的时候手动设计引物就太麻烦了,这里我介绍一个批量查找gap和设计引物的方法:
首先找出GAP的位置:
这里利用一个SSR查找脚本misa,misa是专门查找SSR的(http://pgrc.ipk-gatersleben.de/misa/misa.html),但是我们可以 稍加修改就可以帮我们找到GAP的位置,因为GAP其实是连续的N组成,我们只需要设置misa帮我们查找连续的N就可以了;学过perl正则表达式的人应该会修改的;
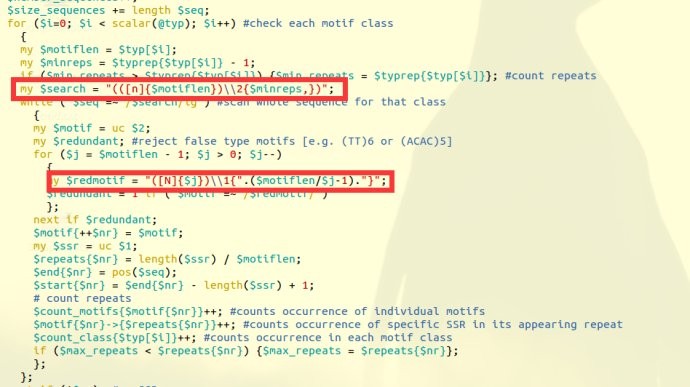
想学习批量设计引物,和正则表达式的应用可以学习一下这个课程:perl高级编程课程
#!/usr/bin/perl -w
use strict;
use Cwd qw(abs_path getcwd);
use Getopt::Long;
use Data::Dumper;
use FindBin qw($Bin $Script);
use File::Basename qw(basename dirname);
use Bio::SeqIO;
my $BEGIN_TIME=time();
my $version="1.0";
# ------------------------------------------------------------------
# GetOptions
# ------------------------------------------------------------------
my ($fa,$flank,$epcrfa,$od,$cut,$outprefix,$PRIMER_NUM_RETURN,$PRIMER_OPT_SIZE,$PRIMER_MIN_SIZE,$PRIMER_MAX_SIZE,$PRIMER_PRODUCT_SIZE_RANGE,$PRIMER_MAX_NS_ACCEPTED);
GetOptions(
"help|?" =>\&USAGE,
"fa=s"=>\$fa,
"flank=s"=>\$flank,
"PRIMER_MIN_SIZE=s"=>\$PRIMER_MIN_SIZE,
"PRIMER_OPT_SIZE=s"=>\$PRIMER_OPT_SIZE,
"PRIMER_MAX_SIZE=s"=>\$PRIMER_MAX_SIZE,
"PRIMER_NUM_RETURN=s"=>\$PRIMER_NUM_RETURN,
"PRIMER_PRODUCT_SIZE_RANGE=s"=>\$PRIMER_PRODUCT_SIZE_RANGE,
"epcrfa=s"=>\$epcrfa,
"od=s"=>\$od,
"prefix=s"=>\$outprefix,
) or &USAGE;
&USAGE unless($fa);
sub USAGE {
my $usage=<<"USAGE";
Program: $0
Version: $version
Contact: huang2002003\@126.com
Discription:
Usage:
Options:
-fa Input fa file, forced
-flank 100 cut 100
-PRIMER_MIN_SIZE 18 Minimum acceptable length of a primer. Must be greater than 0 and less than or equal to
PRIMER_MAX_SIZE
-PRIMER_OPT_SIZE 20 Optimum length (in bases) of a primer. Primer3 will attempt to pick primers close to this length.
-PRIMER_MAX_SIZE 23 Maximum acceptable length (in bases) of a primer. Currently this parameter cannot be
larger than 35. This limit is governed by maximum oligo size for which primer3's melting-temperature
is valid.
-PRIMER_PRODUCT_SIZE_RANGE 100-300 The associated values specify the lengths of the product that the user
wants the primers to create, and is a space separated list of elements of the form
-PRIMER_NUM_RETURN 1 Total num primer to search
-epcrfa run E-PCR to filter multi mapped primers; required;
-od Key of output dir,default cwd;
-prefix defoult demo;
-h Help
USAGE
print $usage;
exit 1;
}
$PRIMER_MIN_SIZE||=18;
$PRIMER_OPT_SIZE||=20;
$PRIMER_MAX_SIZE||=23;
$PRIMER_NUM_RETURN||=1;
$PRIMER_MAX_NS_ACCEPTED||=1;
$PRIMER_PRODUCT_SIZE_RANGE||="100-300";
$od||=getcwd;
$flank||=100;
$od=abs_path($od);
unless(-d $od){ mkdir $od;}
$outprefix||="SSR";
$fa=abs_path($fa);
$epcrfa||=$fa;
$epcrfa=abs_path($epcrfa);
#################################################################
my%ref=();
my%ref_len=();
my$in = Bio::SeqIO->new(-file => "$fa" ,
-format => 'Fasta');
while ( my $seq = $in->next_seq() ) {
$ref{$seq->id}=$seq;
$ref_len{$seq->id}=$seq->length;
}
$in->close();
############find SSR by misa#######################
print "perl /biosoft/MISA/my_misa1.pl $fa $od/$outprefix.misa $od/$outprefix.misa.statistics\n";
system("perl /biosoft/MISA/my_misa1.pl $fa $od/$outprefix.misa $od/$outprefix.misa.statistics");
###########################primer design ###############################
open IN,"$od/$outprefix.misa" or die "$!";
open OUT,">$od/primer3.input" or die "$!";
my%ssr_pos=(); #ssr相关信息
my%SSR=(); # SSR type
while(my$line=){
chomp $line;
my@tmp=split(/\t/,$line);
next if ($tmp[0] eq "ID");
my$ID="$tmp[0]_$tmp[5]_$tmp[6]_$tmp[4]";
$ssr_pos{$ID}="$tmp[0]\t$tmp[5]\t$tmp[6]\t$tmp[4]\t$tmp[2]";
my$seq=&get_target_fa($tmp[0],$tmp[5]-$flank,$tmp[6]+$flank);
#print $seq."\n";
my$tar=0;
if(($tmp[5]-$flank)<=0){
$tar=$tmp[5];
}else{
$tar=$flank;
}
if ($seq){
$SSR{$ID}=$tmp[3];
print OUT "SEQUENCE_ID=$ID\n";
print OUT "SEQUENCE_TEMPLATE=$seq\n";
printf OUT ("SEQUENCE_TARGET=%d,%d\n",$tar+1,$tmp[4]);
print OUT "SPRIMER_TASK=pick_detection_primers\n";
print OUT "PRIMER_PICK_LEFT_PRIMER=1\n";
print OUT "PRIMER_PICK_RIGHT_PRIMER=1\n";
print OUT "PRIMER_NUM_RETURN=$PRIMER_NUM_RETURN\n";
print OUT "PRIMER_MAX_END_STABILITY=250\n";
print OUT "PRIMER_MIN_SIZE=$PRIMER_MIN_SIZE\n";
print OUT "PRIMER_OPT_SIZE=$PRIMER_OPT_SIZE\n";
print OUT "PRIMER_MAX_SIZE=$PRIMER_MAX_SIZE\n";
print OUT "PRIMER_MAX_NS_ACCEPTED=1\n";
print OUT "PRIMER_MIN_GC=40.0\nPRIMER_MAX_GC=60.0\n";
print OUT "PRIMER_PRODUCT_SIZE_RANGE=$PRIMER_PRODUCT_SIZE_RANGE\n";
printf OUT ("SEQUENCE_INTERNAL_EXCLUDED_REGION=%d,%d\n",$tar+1,$tmp[4]);
print OUT "=\n";
}
}
close(OUT);
close(IN);
print "/biosoft/Primer3/primer3-2.3.7/src/primer3_core -p3_settings_file=/biosoft/Primer3/primer3-2.3.7/default_settings.txt -output=$od/$outprefix.primers $od/primer3.input\n";
system("/biosoft/Primer3/primer3-2.3.7/src/primer3_core -p3_settings_file=/biosoft/Primer3/primer3-2.3.7/default_settings.txt -output=$od/$outprefix.primers $od/primer3.input");
###############################e-PCR #####################
my%Pseq=();
$/="=\n";
open IN ,"$od/$outprefix.primers" or die "$!";
open OUT,">$od/$outprefix.primers.xls" or die "$!";
print OUT "#SEQUENCE_ID\tPRIMER_LEFT_SEQUENCE\tPRIMER_RIGHT_SEQUENCE\tPRIMER_PAIR_PRODUCT_SIZE\tPRIMER_LEFT_TM\tPRIMER_RIGHT_TM\tPRIMER_LEFT_GC_PERCENT\tPRIMER_RIGHT_GC_PERCENT\tSSR\tSSR_TYPE\n";
while(my$line=){
chomp$line;
my@field=split(/\n/,$line);
my%primers=&get_hash(@field);
next if ! defined($primers{"PRIMER_PAIR_NUM_RETURNED"}) or $primers{"PRIMER_PAIR_NUM_RETURNED"}==0;
if($primers{"PRIMER_PAIR_NUM_RETURNED"}>=1){
for my$i (0..($primers{"PRIMER_PAIR_NUM_RETURNED"}-1)){
$Pseq{"$primers{SEQUENCE_ID}"}{$i}=[$primers{"PRIMER_LEFT_${i}_SEQUENCE"},$primers{"PRIMER_RIGHT_${i}_SEQUENCE"}];
print OUT join("\t",("$primers{SEQUENCE_ID}_${i}",$primers{"PRIMER_LEFT_${i}_SEQUENCE"},$primers{"PRIMER_RIGHT_${i}_SEQUENCE"},$primers{"PRIMER_PAIR_${i}_PRODUCT_SIZE"},$primers{"PRIMER_LEFT_${i}_TM"},$primers{"PRIMER_RIGHT_${i}_TM"},$primers{"PRIMER_LEFT_${i}_GC_PERCENT"},$primers{"PRIMER_RIGHT_${i}_GC_PERCENT"},$SSR{$primers{"SEQUENCE_ID"}}))."\n";
}
}
}
$/="\n";
close(OUT);
print "/biosoft/e-PCR/Linux-x86_64/e-PCR -w9 -f 1 -m100 $od/$outprefix.primers.xls D=100-400 $epcrfa N=1 G=1 T=3 > $od/$outprefix.epcr.out\n";
system("/biosoft/e-PCR/Linux-x86_64/e-PCR -w9 -f 1 -m100 $od/$outprefix.primers.xls D=100-400 $epcrfa N=1 G=1 T=3 > $od/$outprefix.epcr.out");
######################################################################################
open IN ,"$od/$outprefix.epcr.out" or die "$!";
open OUT,">$od/$outprefix.primers.result.xls" or die "$!";
my %P=();
while(my$line=){
chomp$line;
my@tmp=split(/\t/,$line);
next if $tmp[6]+$tmp[7] >1;
$tmp[5]=~s#/.*$##;
my$ssr_id=$tmp[1];
my$num=0;
($ssr_id,$num)=($ssr_id=~/(.*)_(\d+)$/);
$P{$tmp[1]}{"$tmp[1]_$tmp[3]_$tmp[4]"}=[$tmp[1],$ssr_pos{$ssr_id},$Pseq{$ssr_id}{$num}->[0],$Pseq{$ssr_id}{$num}->[1],$tmp[5],@tmp[6..$#tmp]];
}
close(IN);
print OUT "#GAP_ID\tCHR\tGAP_START\tGAP_END\tGAP_LENGTH\tGAP_TYPE\tPRIMER_LEFT_SEQUENCE\tPRIMER_RIGHT_SEQUENCE\tPRIMER_PAIR_PRODUCT_SIZE/PREDICT_RANGE\tMISM\tGAPS\tPRIMER_LEFT_TM\tPRIMER_RIGHT_TM\tPRIMER_LEFT_GC_PERCENT\tPRIMER_RIGHT_GC_PERCENT\tGAP\n";
open OUT1,">$od/$outprefix.NOprimers.result.xls" or die "$!";
for my $id (sort keys %SSR){
for my$i(0..($PRIMER_NUM_RETURN-1)){
if (exists $P{"${id}_$i"}){
my@ps=(keys %{$P{"${id}_$i"}});
if(@ps ==1){
print OUT join("\t",@{$P{"${id}_$i"}{$ps[0]}})."\n";
}
}else{
print OUT1 "${id} not primers\n";
}
}
}
close(OUT);
close(OUT1);
open OUT,">$od/readme.txt" or die "$!";
my $readme=<<"README";
建议使用editplus打开本文件;
*primers.result.xls
#SEQUENCE_ID-- 引物ID (命名规则:GAP所在来源序列_GAP所在位置start_GAP所在位置end_GAP长度_备用引物编号 ; )
PRIMER_LEFT_SEQUENCE-- 左引物
PRIMER_RIGHT_SEQUENCE-- 右引物
PRIMER_PAIR_PRODUCT_SIZE/PREDICT_RANGE--
MISM-- Total number of mismatches for two primers(ePCR)
GAPS-- Total number of gaps for two primers(ePCR)
PRIMER_LEFT_TM-- 退火温度
PRIMER_RIGHT_TM-- 退火温度
PRIMER_LEFT_GC_PERCENT-- GC含量
PRIMER_RIGHT_GC_PERCENT-- GC含量
GAP-- GAP类型
方法:
2.提取GAP左右及其左右序列,用primer3设计引物[2]
3.用Epcr ,在fasta上电子PCR,去除有多处比较的引物,保证引物扩增的特异性[3]
[1] MISA:Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.).
[2] Primer3: Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3 - new capabilities and interfaces. Nucleic Acids Research 40(15):e115
[3] http://www.ncbi.nlm.nih.gov/tools/epcr/
README
print OUT $readme;
close(OUT);
################################################################################
sub get_hash{
my@info=@_;
my%result=();
for my$i(@info){
next if $i=~/^\s*$/;
my($k,$v)=split(/=/,$i);
$result{$k}=$v;
}
return %result;
}
sub get_target_fa(){
my $id=shift;
my $posU=shift;
my $posD=shift;
if($posU<=0){
$posU=1;
}
my $seqobj=$ref{$id};
my $len=$ref_len{$id};
return $seqobj->subseq($posU,$len) if $posD>$len;
my $tri="";
$tri=$seqobj->subseq($posU,$posD);
return $tri;
}
- 发表于 2018-06-15 11:21
- 阅读 ( 4136 )
- 分类:基因组学
你可能感兴趣的文章
相关问题
0 条评论
请先 登录 后评论
